-Search query
-Search result
Showing all 40 items for (author: liu & yh)
EMDB-39012: 
Representative tomogram of primary glioblastoma stem cell with circular inter-mitochondrial junctions.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
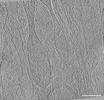

EMDB-39015: 
Representative tomogram of microglia cell with nanotunnel-like structures resembling mitochondrial fission.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
EMDB-39019: 
Representative tomogram of glioblastoma cell with nanotunnel-like structure and inter-mitochondrial junction.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-39021: 
Representative tomogram of normal human astrocyte with nanotunnel-like structure which is an extension of the mitochondrial outer membrane.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
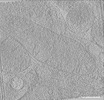

EMDB-39023: 
Representative tomogram of primary glioblastoma differentiated cell with parallel inter-mitochondrial junction.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT
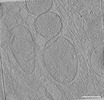

EMDB-39024: 
Representative tomogram of primary glioblastoma stem cell with clustered mitochondria bearing various long-short axis ratios.
Method: electron tomography / : Wang R, Lei H, Wang HX, Qi L, Liu YE, Liu YH, Shi YF, Chen JX, Shen QT

EMDB-33145: 
Cryo-EM structures of human mitochondrial NAD(P)+-dependent malic enzyme in apo form
Method: single particle / : Wang CH, Hsieh JT, Ho MC, Hung HC

EMDB-33146: 
Cryo-EM structures of human mitochondrial NAD(P)+-dependent malic enzyme in a ternary complex with NAD+ and allosteric inhibitor EA
Method: single particle / : Wang CH, Hsieh JT, Ho MC, Hung HC

EMDB-33147: 
Cryo-EM structures of human mitochondrial NAD(P)+-dependent malic enzyme in a ternary complex with NAD+ and allosteric inhibitor MDSA
Method: single particle / : Wang CH, Hsieh JT, Ho MC, Hung HC

EMDB-35522: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex(mask on receptor)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35523: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex(mask on Giq-scFV16 complex)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35524: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi1 complex(mask on receptor)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35525: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi1 complex(mask on Gil-scFV16 complex)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35529: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex (consensus map)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35533: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi complex(consensus map)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35356: 
Cryo-EM structure of the 9-hydroxystearic acid bound GPR120-Gi complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35357: 
Cryo-EM structure of the linoleic acid bound GPR120-Gi complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35358: 
Cryo-EM structure of the oleic acid bound GPR120-Gi complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35359: 
Cryo-EM structure of the TUG891 bound GPR120-Gi complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35360: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-29736: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-32928: 
Cryo-EM Structure of Arabidopsis CRY2 in active conformation
Method: single particle / : Hao YH, Zhang X, Zhang P

EMDB-32929: 
Cryo-EM Structure of Arabidopsis CRY2 tetramer in complex with CIB1 fragment
Method: single particle / : Hao YH, Zhang X, Zhang P

EMDB-32832: 
SARS-CoV-2 Spike in complex with Fab of m31A7
Method: single particle / : Wu YM, Chen X

EMDB-32328: 
Cryo-EM structure of GmALMT12/QUAC1 anion channel
Method: single particle / : Qin L, Tang LH, Xu JS, Zhang XH, Zhu Y, Sun F, Su M, Zhai YJ, Chen YH

EMDB-32825: 
Negative stain volume of the mono-GlcNAc-decorated SARS-CoV-2 Spike
Method: single particle / : Chen X, Huang HY

EMDB-31470: 
Cryo-EM structure of SARS-CoV-2 spike in complex with a neutralizing antibody chAb-25 (Focused refinement of S-RBD and chAb-25 region)
Method: single particle / : Yang TJ, Yu PY, Wu HC, Hsu STD

EMDB-31471: 
Cryo-EM structure of SARS-CoV-2 spike in complex with a neutralizing antibody chAb-45 (Focused refinement of S-RBD and chAb-45 region)
Method: single particle / : Yang TJ, Yu PY, Wu HC, Hsu STD

EMDB-30392: 
Cryo-EM structure of Fenoldopam bound dopamine receptor DRD1-Gs signaling complex
Method: single particle / : Yan W, Shao W

EMDB-30393: 
Cryo-EM structure of A77636 bound dopamine receptor DRD1-Gs signaling complex
Method: single particle / : Yan W, Shao Z

EMDB-30394: 
Cryo-EM structure of PW0464 bound dopamine receptor DRD1-Gs signaling complex
Method: single particle / : Yan W, Shao Z

EMDB-30395: 
Cryo-EM structure of Dopamine and LY3154207 bound dopamine receptor DRD1-Gs signaling complex
Method: single particle / : Yan W, Shao Z

EMDB-30452: 
Cryo-EM structure of SKF83959 bound dopamine receptor DRD1-Gs signaling complex
Method: single particle / : Yan W, Shao ZH

EMDB-10462: 
Leishmania tarentolae proteasome 20S subunit complexed with LXE408
Method: single particle / : Srinivas H

EMDB-10463: 
Leishmania tarentolae proteasome 20S subunit complexed with LXE408 and bortezomib
Method: single particle / : Srinivas H

EMDB-9790: 
Cryo-EM structure and transport mechanism of a wall teichoic acid ABC transporter
Method: single particle / : Chen L, Hou WT

EMDB-6976: 
Structure of the Herpes simplex virus type 2 C-capsid with capsid-vertex-specific component
Method: single particle / : Wang JL, Yuan S, Zhu DJ, Tang H, Wang N, Chen WY, Gao Q, Li YH, Wang JZ, Liu HR, Zhang XZ, Rao ZH, Wang XX

EMDB-8629: 
Conformational states of a soluble, uncleaved HIV-1 envelope trimer
Method: single particle / : Liu YH, Pan JH, Cai YF, Grigorieff N, Harrison SC, Chen B
EMDB-8631: 
Conformational states of a soluble, uncleaved HIV-1 envelope trimer
Method: single particle / : Liu YH, Pan JH, Cai YF, Grigorieff N, Harrison SC, Chen B
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
